Note
Go to the end to download the full example code.
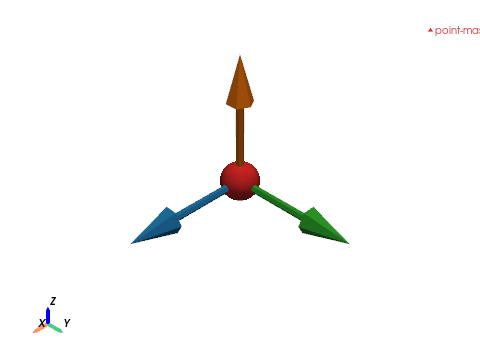
POINT_MASS — single-node lumped mass#
The 1-node concentrated-mass element contributes a diagonal
3 x 3 block to the global mass matrix:
where \((m_x, m_y, m_z)\) are the per-axis masses (typically all equal to a single isotropic mass \(m\)). Consistent and lumped formulations coincide for a one-node element.
The rotary-inertia variant (KEYOPT(3) != 2) adds three
diagonal rotary-inertia entries; that path is signposted in
the kernel module header as a future addition.
References#
Cook, R. D., Malkus, D. S., Plesha, M. E., Witt, R. J. (2002) Concepts and Applications of Finite Element Analysis, 4th ed., Wiley, §11.3.
Bathe, K.-J. (2014) Finite Element Procedures, 2nd ed., Prentice Hall, §4.2.2.
Implementation: femorph_solver.elements.point_mass.PointMass.
from __future__ import annotations
import numpy as np
import pyvista as pv
Render the single-node element#
origin = np.array([[0.0, 0.0, 0.0]])
plotter = pv.Plotter(off_screen=True, window_size=(480, 360))
plotter.add_points(
origin, render_points_as_spheres=True, point_size=40, color="#d62728", label="point-mass node"
)
# Three axis-aligned arrows showing the 3 mass DOFs (m_x, m_y, m_z).
for vec, _label, col in [
(np.array([1.0, 0.0, 0.0]), "m_x", "#1f77b4"),
(np.array([0.0, 1.0, 0.0]), "m_y", "#2ca02c"),
(np.array([0.0, 0.0, 1.0]), "m_z", "#ff7f0e"),
]:
plotter.add_arrows(origin, vec[None, :], mag=0.8, color=col)
plotter.add_axes(line_width=4, color="black")
plotter.view_isometric()
plotter.camera.zoom(0.9)
plotter.add_legend(face=None, size=(0.22, 0.07), bcolor="white")
plotter.show()
Sanity — global mass contribution#
A point-mass at node n adds M_e to rows / columns
(3n, 3n+1, 3n+2) of the global mass matrix. Verify the
contribution is exactly diagonal.
point-mass contribution at one node:
M_e (diag) = [7.5 7.5 7.5]
OK — M_e is purely diagonal.
Total running time of the script: (0 minutes 0.145 seconds)